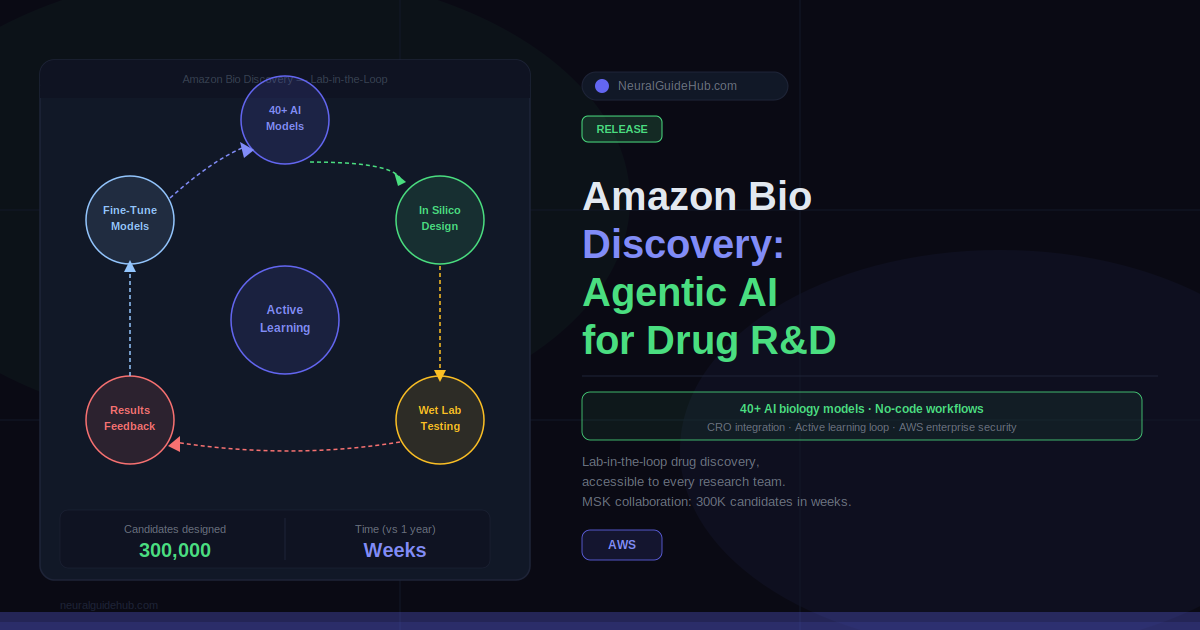
Drug discovery has a collaboration problem that the field has been quietly tolerating for years. Computational biologists have the models. Bench scientists have the biological intuition. But getting them to work from the same system, on the same data, without a queue of manual handoffs between them — that has been the unsolved part.
Amazon Bio Discovery is AWS’s answer to that. Launched as a new agentic application, it brings computational design and wet-lab validation together in a single platform. The goal is to make lab-in-the-loop drug discovery accessible not just to organizations with dedicated infrastructure teams, but to any research organization that needs it.
The numbers are not abstract. In a collaboration with Memorial Sloan Kettering Cancer Center, Amazon Bio Discovery was used to design nearly 300,000 novel antibody candidates, filter to the top 100,000, and send them to wet-lab testing. Time required: weeks. The traditional approach for comparable work: up to a year.
The Problem Amazon Bio Discovery Is Built to Solve
Most research organizations running AI-assisted drug discovery face the same structural problem. New biological AI models emerge constantly, each with different strengths and integration requirements. Computational biologists are expected to evaluate, operationalize, and maintain these models across a growing number of programs, without the infrastructure to match the pace.
Bench scientists, on the other hand, have deep expertise in their targets and experiments but no direct path to the computational tools that could accelerate their work. The result is a bottleneck at the handoff. Not because the science is unavailable, but because the tooling does not support how these teams actually need to work together.
When computational predictions and wet-lab workflows run in separate systems, manual handoffs introduce delays, reproducibility suffers, and the feedback loop that makes lab-in-the-loop valuable in the first place slows to a crawl.

What Amazon Bio Discovery Actually Provides
The platform provides access to 40+ AI biology models with AI-guided selection. Users can also upload custom models and models licensed from third parties. Agentic assistants help select the right models for specific research goals, optimize configurations, and evaluate candidates for experimentation.
For computational biologists, this means building and modifying workflows in a no-code environment without managing infrastructure or provisioning compute for training and inference. Workflows can be published for team-wide use, with standardized data processing and rigorous analysis built in.
For bench scientists, it means running multiple experiment versions in parallel and adjusting parameters through agentic assistance, rather than waiting for a custom solution to be built. Both roles work from the same system, the same data, and the same results.
Amazon Bio Discovery’s CRO partners — Ginkgo Bioworks, Twist Bioscience, and A-Alpha Bio — enable direct wet-lab validation with results flowing back automatically to improve the next cycle.
The 5-Step Workflow in Practice
Step 1: Evaluate Models and Build Your In Silico Workflow
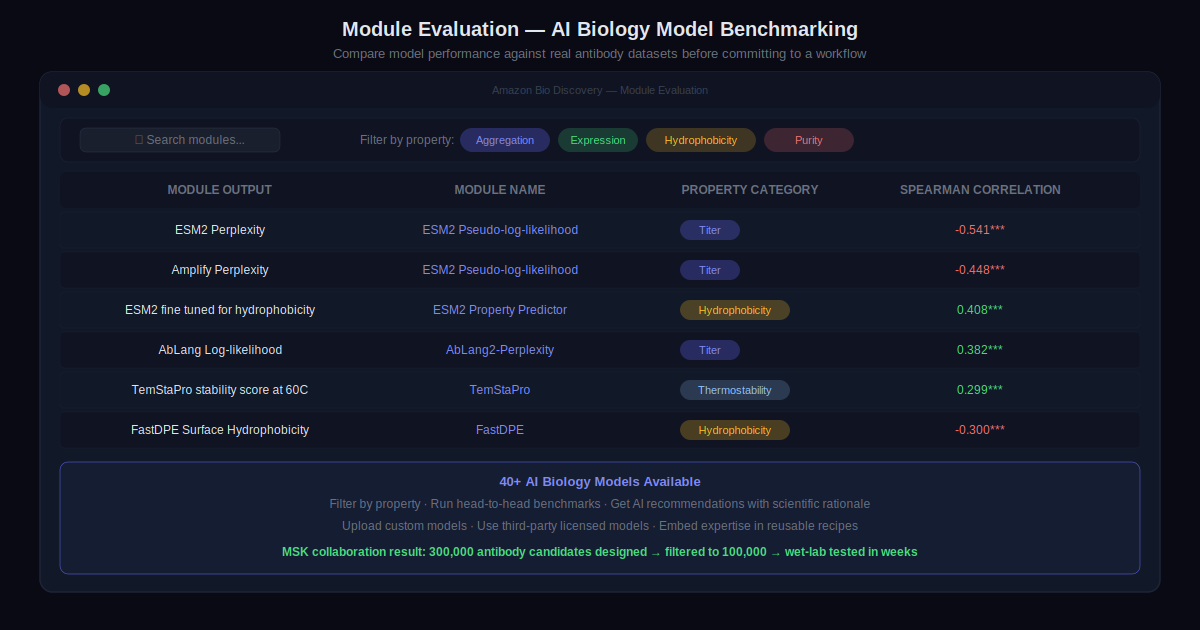
The model catalog covers 40+ specialized AI biology models. The module evaluation page lets users filter by target property and run head-to-head comparisons using antibody developability benchmarks. The AI-assisted workflow walks through objectives and generates a list of model recommendations with scientific rationale.
The output is a recipe — a computational workflow pipeline that can be modified in a sandbox and published for team use. Computational expertise gets embedded in the recipe. Biological expertise drives how it is applied.
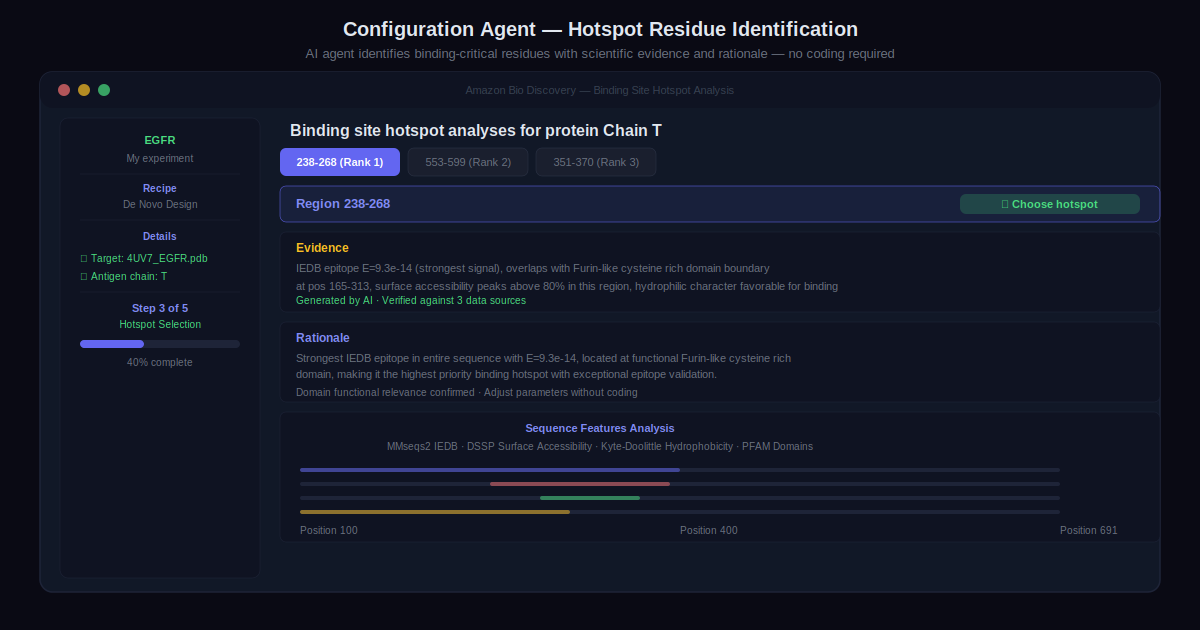
Step 2: Design Your Experiment with Configuration Agents
The AI agent guides experiment setup. For antibody design, that means identifying hotspot residues and selecting the structural framework that affects antigen binding. The agent searches multiple data sources, considers surface accessibility and hydrophobicity, and provides recommendations with scientific rationale and references.
Parameters are adjustable directly through the interface. No coding required at any point in this process.
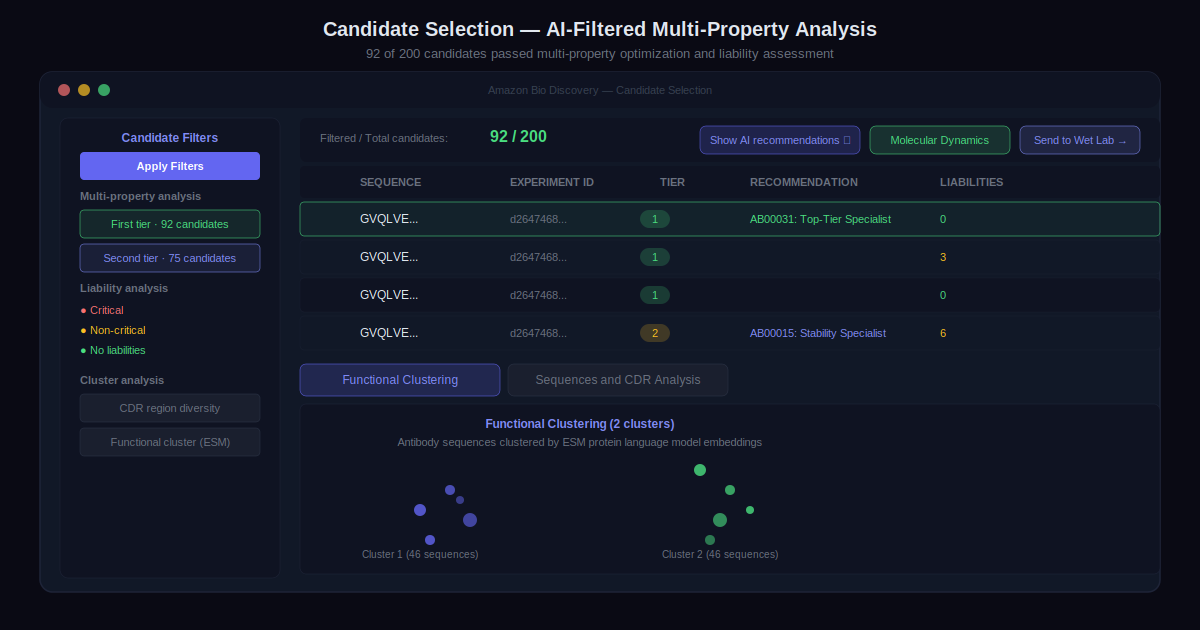
Step 3: Analyze Results with Candidate Selection Agents
When the in silico experiment completes, an AI-generated summary and a pre-filtered candidate list appear automatically. Candidates have already been run through multi-property optimization and liability assessment, checking for chemical modifications that could impact antibody stability, efficacy, or developability.
Each recommended candidate comes with rationale. Advanced analytical tools — molecular dynamics analysis, functional and sequential diversity reviews — are available for deeper evaluation before deciding which candidates to advance.
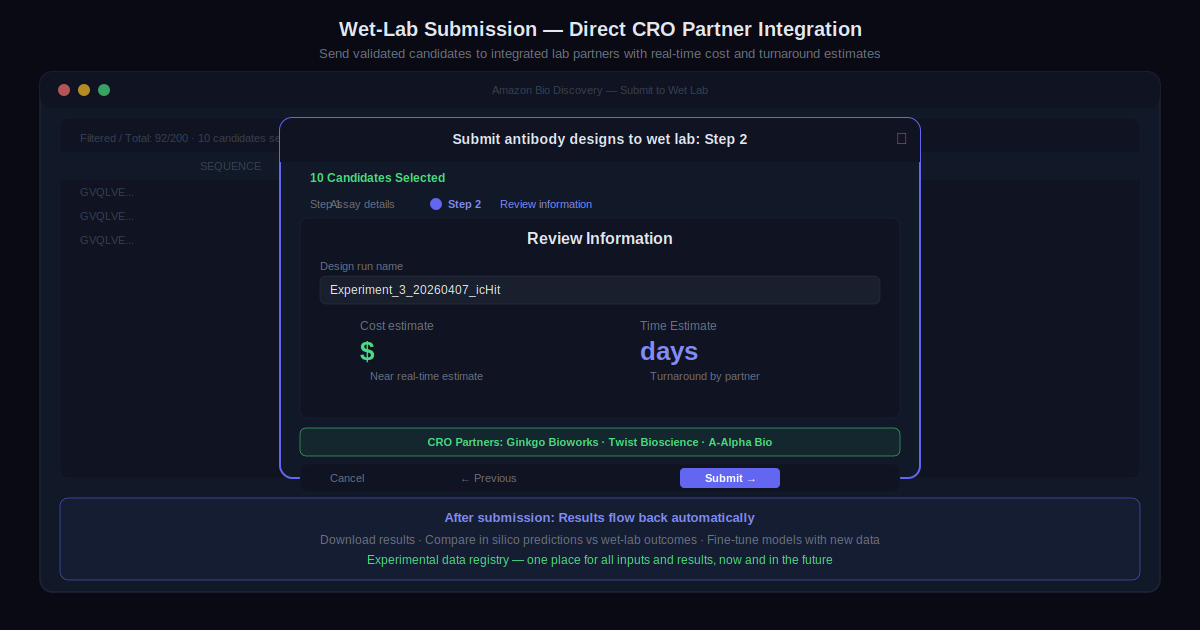
Step 4: Send Top Candidates to the Wet Lab
Validated candidates go directly to one of the integrated CRO partners. Users select assays and receive near-real-time cost and turnaround estimates. No manual handoffs. No custom integrations with each partner’s systems.
When testing completes, results flow back into Amazon Bio Discovery automatically. In silico predictions can be compared against wet-lab outcomes directly, and the experimental data registry provides a single view of all inputs and results over time.
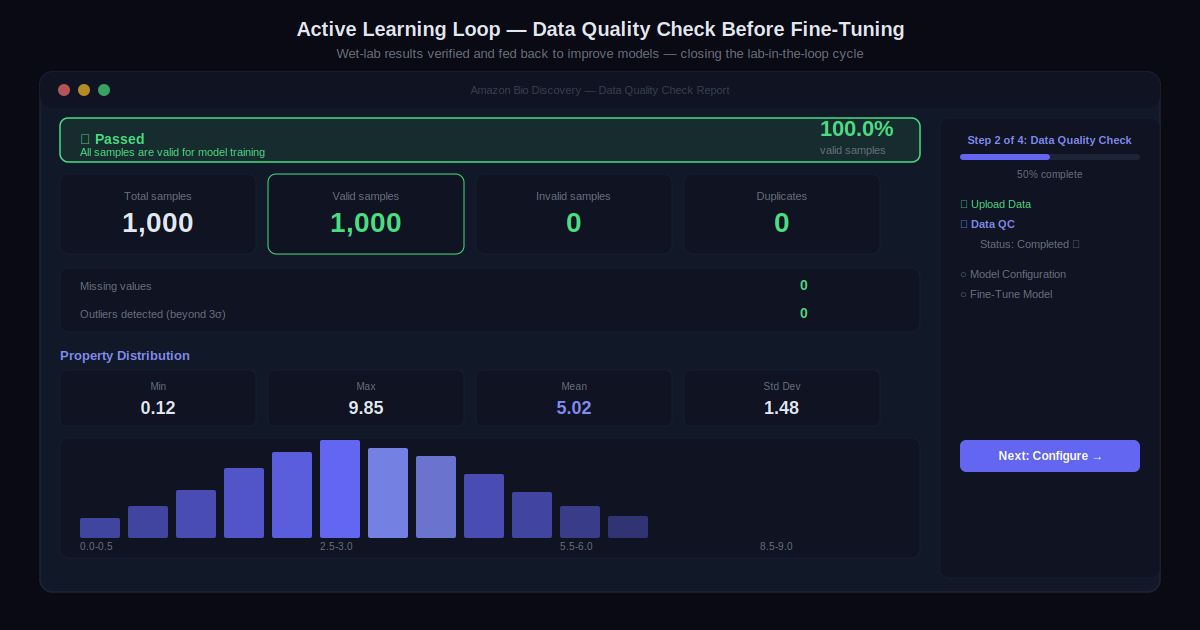
Step 5: Close the Loop with Active Learning
Wet-lab results feed back into the system for model fine-tuning. After a few cycles, the system identifies drug-like candidates with increasing confidence. The models improve with each iteration, and both the computational workflows and the biological predictions become more accurate over time.
Infrastructure and Enterprise Access
Amazon Bio Discovery runs on the same AWS infrastructure used by 19 of the top 20 pharmaceutical companies. Data isolation ensures proprietary experimental data and custom-trained models remain protected within each application environment.
The collaboration with Memorial Sloan Kettering remains the clearest demonstration of what this looks like at scale: 300,000 antibody candidates designed, filtered to 100,000, and sent to wet-lab testing in weeks. The traditional timeline for comparable work runs up to a year.
For research organizations that have been running computational and wet-lab workflows in separate systems with manual handoffs between them, the consolidation alone is a significant operational change. The active learning loop on top of that is what makes the platform worth watching as it scales across more programs and research teams.
https://aws.amazon.com/blogs/industries/introducing-amazon-bio-discovery